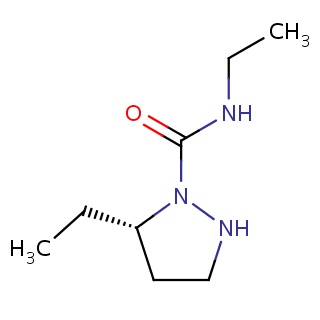
Binding information for 2g1q_ligand_3_16.mol2(FDBF02647)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 2g1q_ligand_3_16.mol2 | 2g1q | 0.596774 | -6.23 | C(C)NC(=O)N1NCC[C@@H]1CC | 12 |
Structure and binding mode of 2g1q_ligand_3_16.mol2(FDBF02647)
Important binding residues for 2g1q_ligand_3_16.mol2(FDBF02647)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 2g1q | GLU118 | -0.62 | 0.21 | -0.41 | 0.07 | -0.35 |
| 2g1q | ARG119 | -1.13 | -5.37 | -6.5 | 4.94 | -1.56 |
| 2g1q | PRO137 | -0.45 | 0.06 | -0.39 | 0.00 | -0.38 |
| 2g1q | TYR211 | -1.27 | 0.18 | -1.09 | 0.67 | -0.42 |
| 2g1q | LEU214 | -1.38 | -0.84 | -2.22 | 0.68 | -1.53 |
| 2g1q | GLU215 | -0.98 | -0.16 | -1.14 | 0.79 | -0.35 |
| 2g1q | ALA218 | -0.78 | 0.35 | -0.43 | -0.52 | -0.96 |