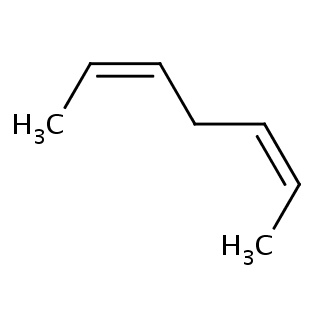
Binding information for 1gnj_ligand_4_680.mol2(FDBF00083)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 1gnj_ligand_4_680.mol2 | 1gnj | 1 | -6.81 | C/C=C\C/C=C\C | 7 |
Structure and binding mode of 1gnj_ligand_4_680.mol2(FDBF00083)
Important binding residues for 1gnj_ligand_4_680.mol2(FDBF00083)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 1gnj | TYR138 | -0.75 | 0.11 | -0.64 | -0.03 | -0.67 |
| 1gnj | LEU139 | -1.04 | 0.06 | -0.98 | -0.03 | -1.00 |
| 1gnj | ILE142 | -1.33 | -0.11 | -1.44 | -0.25 | -1.68 |
| 1gnj | LEU154 | -0.81 | -0.06 | -0.87 | 0.26 | -0.61 |
| 1gnj | PHE157 | -1.09 | -0.09 | -1.18 | 0.06 | -1.12 |
| 1gnj | ALA158 | -1.01 | -0.05 | -1.06 | 0.05 | -1.02 |
| 1gnj | TYR161 | -1.44 | -0.23 | -1.67 | 0.60 | -1.08 |