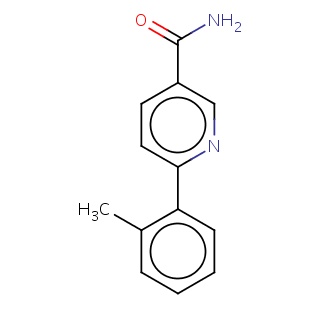
Binding information for 3iph_ligand_2_1.mol2(FDBF02971)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 3iph_ligand_2_1.mol2 | 3iph | 0.503937 | -8.33 | c1(ccccc1C)c1ccc(cn1)C(=O)N | 16 |
Structure and binding mode of 3iph_ligand_2_1.mol2(FDBF02971)
Important binding residues for 3iph_ligand_2_1.mol2(FDBF02971)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 3iph | VAL38 | -0.92 | -0.05 | -0.97 | 0.01 | -0.96 |
| 3iph | ALA51 | -1.31 | -0.14 | -1.45 | 0.26 | -1.20 |
| 3iph | LYS53 | -1.65 | -1.39 | -3.04 | 1.32 | -1.72 |
| 3iph | ILE84 | -0.92 | 0.06 | -0.86 | -0.11 | -0.96 |
| 3iph | LEU104 | -0.86 | -0.17 | -1.03 | 0.12 | -0.90 |
| 3iph | VAL105 | -0.28 | 0.02 | -0.26 | -0.08 | -0.34 |
| 3iph | LEU108 | -1.27 | -1.48 | -2.75 | 0.60 | -2.15 |
| 3iph | MET109 | -0.48 | -2.85 | -3.33 | 1.29 | -2.04 |
| 3iph | ALA157 | -0.37 | -0.38 | -0.75 | 0.23 | -0.52 |
| 3iph | LEU167 | -1.85 | 0.13 | -1.72 | -0.15 | -1.87 |