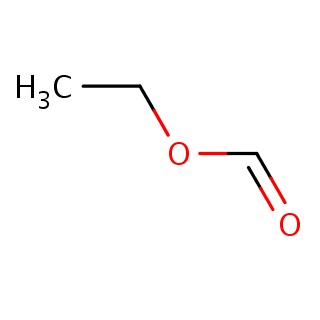
Binding information for 1p6d_ligand_1_4.mol2(FDBF00100)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 1p6d_ligand_1_4.mol2 | 1p6d | 1 | -5.51 | C(OC=O)C | 5 |
Structure and binding mode of 1p6d_ligand_1_4.mol2(FDBF00100)
Important binding residues for 1p6d_ligand_1_4.mol2(FDBF00100)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 1p6d | PHE66 | -0.96 | 0.06 | -0.9 | -0.02 | -0.92 |
| 1p6d | PHE70 | -0.73 | -0.34 | -1.07 | 0.21 | -0.87 |
| 1p6d | ASP122 | -0.04 | -0.76 | -0.8 | -0.10 | -0.89 |
| 1p6d | THR133 | -0.63 | -1.60 | -2.23 | 0.57 | -1.65 |
| 1p6d | ASN134 | -0.38 | -2.22 | -2.6 | 1.67 | -0.93 |
| 1p6d | GLU146 | -0.21 | -0.91 | -1.12 | -1.81 | -2.94 |