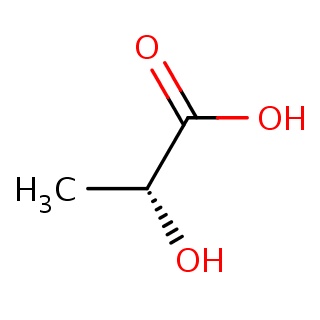
Binding information for 1nf8_ligand_1_1.mol2(FDBF03150)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 1nf8_ligand_1_1.mol2 | 1nf8 | 1 | -6.20 | O[C@@H](C(=O)O)C | 6 |
Structure and binding mode of 1nf8_ligand_1_1.mol2(FDBF03150)
Important binding residues for 1nf8_ligand_1_1.mol2(FDBF03150)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 1nf8 | HIS37 | -0.71 | -20.65 | -21.36 | 20.59 | -0.78 |
| 1nf8 | LYS122 | 0.35 | -63.72 | -63.37 | 56.63 | -6.75 |
| 1nf8 | ARG124 | -0.11 | -13.35 | -13.46 | 13.00 | -0.46 |
| 1nf8 | TYR125 | -0.65 | 1.57 | 0.92 | -1.85 | -0.94 |
| 1nf8 | GLY149 | -0.02 | -18.35 | -18.37 | 18.06 | -0.32 |
| 1nf8 | VAL150 | -0.82 | 0.32 | -0.5 | -0.96 | -1.46 |
| 1nf8 | TYR151 | -1.26 | -1.16 | -2.42 | -0.03 | -2.46 |
| 1nf8 | VAL154 | -0.94 | -5.94 | -6.88 | 2.39 | -4.49 |
| 1nf8 | GLY155 | 0.24 | -8.99 | -8.75 | 4.49 | -4.26 |