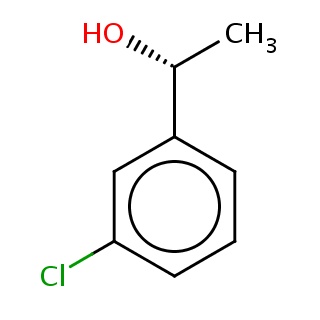
Binding information for 3gw5_ligand_1_0.mol2(FDBF03178)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 3gw5_ligand_1_0.mol2 | 3gw5 | 0.697674 | -6.90 | [C@H](O)(c1cccc(c1)Cl)C | 10 |
Structure and binding mode of 3gw5_ligand_1_0.mol2(FDBF03178)
Important binding residues for 3gw5_ligand_1_0.mol2(FDBF03178)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 3gw5 | GLN19 | -1.35 | 0.36 | -0.99 | -0.01 | -0.99 |
| 3gw5 | PRO118 | -0.79 | -0.84 | -1.63 | 0.65 | -0.98 |
| 3gw5 | PHE119 | -0.94 | 0.12 | -0.82 | 0.19 | -0.62 |
| 3gw5 | LEU121 | -1.02 | 0.10 | -0.92 | 0.22 | -0.70 |
| 3gw5 | ALA122 | -0.94 | -0.46 | -1.4 | 0.15 | -1.24 |
| 3gw5 | PHE124 | -0.83 | -0.15 | -0.98 | 0.35 | -0.63 |
| 3gw5 | SER230 | 1.50 | -4.14 | -2.64 | 1.89 | -0.75 |