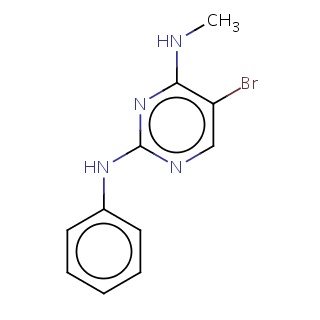
Binding information for 1z5m_ligand_4_1.mol2(FDBF03488)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 1z5m_ligand_4_1.mol2 | 1z5m | 0.753425 | -7.93 | c1(ccccc1)Nc1nc(c(cn1)Br)NC | 16 |
Structure and binding mode of 1z5m_ligand_4_1.mol2(FDBF03488)
Important binding residues for 1z5m_ligand_4_1.mol2(FDBF03488)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 1z5m | LEU88 | -3.34 | -0.99 | -4.33 | 1.37 | -2.95 |
| 1z5m | GLY89 | -0.51 | 0.35 | -0.16 | -0.21 | -0.37 |
| 1z5m | VAL96 | -1.42 | -0.23 | -1.65 | -0.19 | -1.84 |
| 1z5m | ALA109 | -1.14 | -0.14 | -1.28 | 0.06 | -1.22 |
| 1z5m | VAL143 | -0.68 | -0.27 | -0.95 | 0.46 | -0.48 |
| 1z5m | TYR161 | -1.71 | -1.23 | -2.94 | 1.06 | -1.88 |
| 1z5m | ALA162 | 0.71 | -4.99 | -4.28 | 1.66 | -2.62 |
| 1z5m | LYS163 | -0.82 | -0.33 | -1.15 | 0.70 | -0.44 |
| 1z5m | ASN164 | -0.45 | -0.16 | -0.61 | 0.01 | -0.59 |
| 1z5m | GLY165 | -1.06 | -0.51 | -1.57 | 0.25 | -1.32 |
| 1z5m | LEU212 | -1.72 | -0.10 | -1.82 | -0.00 | -1.82 |
| 1z5m | THR222 | -0.66 | 0.10 | -0.56 | 0.06 | -0.50 |