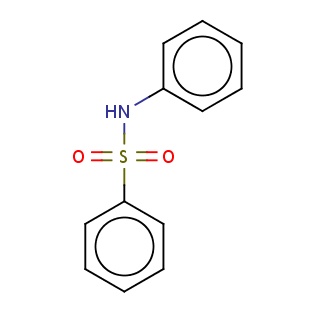
Binding information for 7upj_ligand_2_9.mol2(FDBF03804)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 7upj_ligand_2_9.mol2 | 7upj | 0.712121 | -7.01 | N(S(=O)(=O)c1ccccc1)c1ccccc1 | 16 |
Structure and binding mode of 7upj_ligand_2_9.mol2(FDBF03804)
Important binding residues for 7upj_ligand_2_9.mol2(FDBF03804)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 7upj | GLY27 | -0.58 | -0.93 | -1.51 | 1.18 | -0.32 |
| 7upj | ALA28 | -1.40 | -1.53 | -2.93 | 0.59 | -2.34 |
| 7upj | ASP29 | -1.76 | 0.76 | -1 | 0.01 | -1.00 |
| 7upj | ILE47 | -0.90 | -0.24 | -1.14 | -0.01 | -1.16 |
| 7upj | GLY48 | -0.84 | -3.67 | -4.51 | 2.93 | -1.58 |
| 7upj | GLY49 | -0.54 | -0.08 | -0.62 | 0.15 | -0.47 |
| 7upj | ILE84 | -0.44 | 0.12 | -0.32 | -0.13 | -0.45 |
| 7upj | ARG8 | -1.53 | -0.49 | -2.02 | 0.78 | -1.25 |
| 7upj | ILE50 | -0.88 | 0.01 | -0.87 | -0.12 | -0.99 |
| 7upj | PRO81 | -0.32 | 0.22 | -0.1 | -0.23 | -0.32 |
| 7upj | VAL82 | -0.44 | -0.04 | -0.48 | 0.06 | -0.43 |