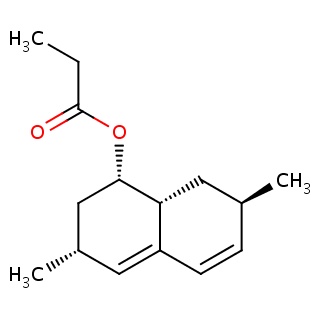
Binding information for 1xdd_ligand_1_1.mol2(FDBF00148)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 1xdd_ligand_1_1.mol2 | 1xdd | 0.84 | -7.98 | [C@@H]1(C[C@H](C=C2[C@H]1C[C@H](C=C2)C)C)OC(=O)CC | 17 |
Structure and binding mode of 1xdd_ligand_1_1.mol2(FDBF00148)
Important binding residues for 1xdd_ligand_1_1.mol2(FDBF00148)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 1xdd | LEU132 | -0.66 | 0.01 | -0.65 | -0.04 | -0.68 |
| 1xdd | VAL157 | -0.68 | 0.63 | -0.05 | -0.36 | -0.41 |
| 1xdd | VAL233 | -0.48 | 0.01 | -0.47 | -0.11 | -0.58 |
| 1xdd | ILE235 | -1.23 | 0.01 | -1.22 | -0.13 | -1.35 |
| 1xdd | TYR257 | -2.55 | -0.99 | -3.54 | 1.33 | -2.21 |
| 1xdd | LEU298 | -1.10 | -0.16 | -1.26 | 0.25 | -1.01 |
| 1xdd | LEU302 | -2.75 | -0.05 | -2.8 | -0.12 | -2.92 |
| 1xdd | ILE306 | -0.53 | -0.02 | -0.55 | -0.07 | -0.61 |