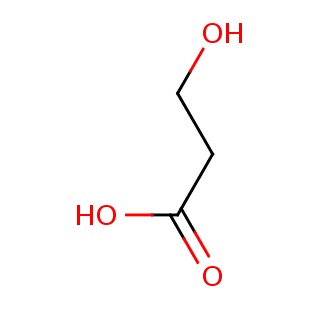
Binding information for 3cda_ligand_1_8.mol2(FDBF00149)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 3cda_ligand_1_8.mol2 | 3cda | 1 | -5.89 | C(O)CC(=O)O | 6 |
Structure and binding mode of 3cda_ligand_1_8.mol2(FDBF00149)
Important binding residues for 3cda_ligand_1_8.mol2(FDBF00149)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 3cda | ARG590 | -0.33 | -45.01 | -45.34 | 39.13 | -6.21 |
| 3cda | MET657 | -0.32 | -14.93 | -15.25 | 14.75 | -0.50 |
| 3cda | SER684 | 0.15 | -9.68 | -9.53 | 8.87 | -0.66 |
| 3cda | THR689 | -0.08 | -1.92 | -2 | 1.68 | -0.32 |
| 3cda | ASP690 | -0.22 | 18.95 | 18.73 | -20.54 | -1.80 |
| 3cda | LYS691 | -0.41 | -17.24 | -17.65 | 17.06 | -0.59 |
| 3cda | LYS692 | -0.60 | -24.07 | -24.67 | 21.99 | -2.67 |
| 3cda | LYS735 | 0.71 | -72.11 | -71.4 | 63.71 | -7.69 |
| 3cda | ALA751 | -0.39 | 2.32 | 1.93 | -2.55 | -0.61 |
| 3cda | LEU853 | -0.69 | -0.74 | -1.43 | 0.36 | -1.07 |
| 3cda | LEU857 | -0.27 | -1.14 | -1.41 | 1.06 | -0.36 |