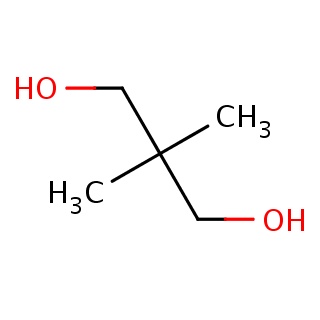
Binding information for 3bex_ligand_2_0.mol2(FDBF00151)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 3bex_ligand_2_0.mol2 | 3bex | 1 | -6.22 | C(C)(C)(CO)CO | 7 |
Structure and binding mode of 3bex_ligand_2_0.mol2(FDBF00151)
Important binding residues for 3bex_ligand_2_0.mol2(FDBF00151)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 3bex | VAL62 | -0.89 | 0.09 | -0.8 | -0.18 | -0.98 |
| 3bex | VAL102 | -0.09 | -0.26 | -0.35 | -0.09 | -0.44 |
| 3bex | GLY103 | -0.23 | -0.32 | -0.55 | 0.03 | -0.51 |
| 3bex | ASP105 | 1.26 | -18.20 | -16.94 | 15.21 | -1.73 |
| 3bex | ARG106 | -0.89 | -1.22 | -2.11 | 0.36 | -1.75 |
| 3bex | ALA129 | -0.71 | -0.09 | -0.8 | 0.15 | -0.64 |
| 3bex | THR131 | -0.71 | -0.06 | -0.77 | 0.25 | -0.52 |
| 3bex | ILE145 | -0.78 | 0.01 | -0.77 | -0.06 | -0.84 |
| 3bex | THR160 | -0.51 | -0.10 | -0.61 | 0.18 | -0.44 |
| 3bex | LEU163 | -0.79 | -0.04 | -0.83 | -0.03 | -0.86 |